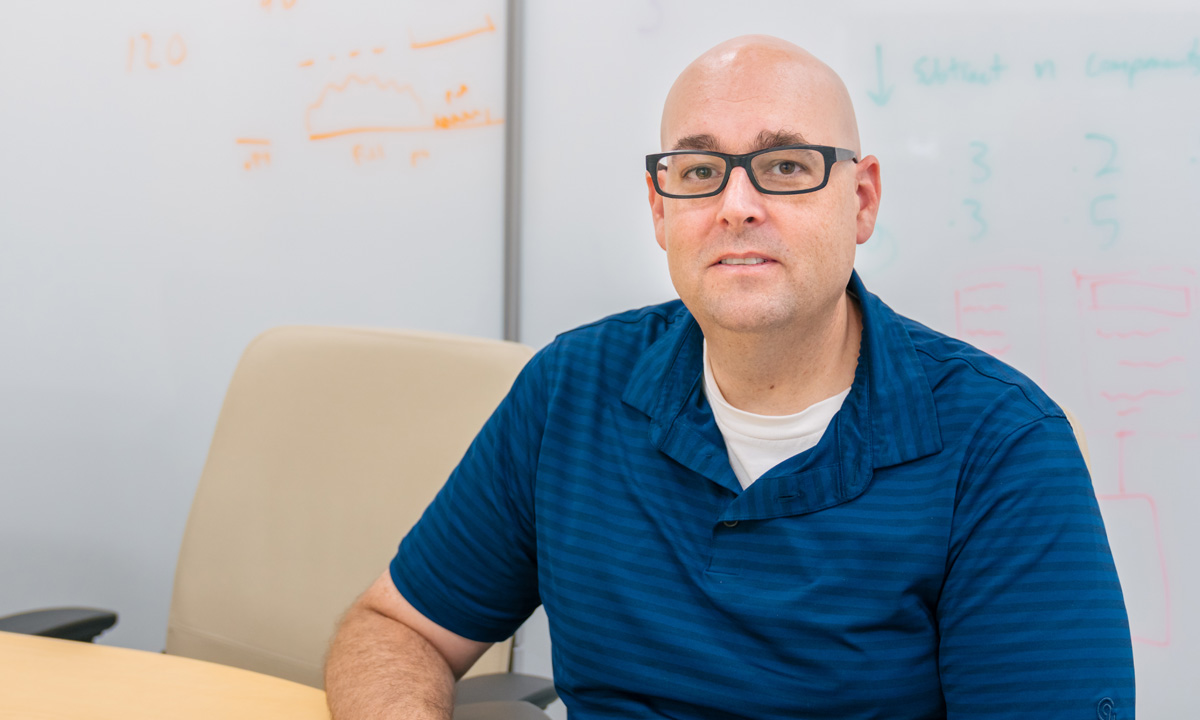
Keith Simmon is part of ARUP’s 15-member bioinformatics team, which supports next-generation sequencing testing.
Keith Simmon returned to ARUP in March 2014 to join the bioinformatics team and shortly after received his PhD in biomedical informatics. Recently, he was selected to speak at the Association for Molecular Pathology (AMP) annual conference this November in Salt Lake City.
“Of all the abstracts submitted to the AMP for consideration in the Informatics category, only four abstracts are accepted by the program committee for an oral informatics platform presentation each year,” points out a proud Elaine Gee, who is the director of bioinformatics at ARUP.
Keith’s presentation is titled: “Homopolymer Compression Improves Reference-Free, Kmer Based Whole Genome Strain Comparison for IonTorrent Data.” A topic that can only be truly appreciated by others in the bioinformatics field.
Keith’s background was originally in microbiology and from 11/29/2004 to 03/18/2011 he worked for ARUP as an R&D scientist on infectious diseases. He helped with the development of Taxonomer, a DNA search engine that rapidly identifies any microorganism by its genetic material.
In his presentation, Keith will focus on how to determine if two or more bacterial strains are part of the same outbreak. “To do this, we sequence the genome of each strain and then compare the genome sequences. The comparison is the difficult part,” explains Keith.
Instead of trying to assemble the sequence pieces into long genome fragment, the task is simplified by looking at the distribution of “words”—in bioinformatics lingo, these are “’kmers.” “This leaves us with a word list for each bacterial strain. Then, we can simply look at how many words are shared to get an idea if the strains are related.”
The ARUP bioinformatics group was formed in 2014 and has grown to include 15 members to support next-generation sequencing testing. Clinical bioinformatics is an exploding field that requires strong proficiency in biology, computer science (i.e., programming, algorithms, big data), data analysis, and visualization to create new analytical tools to advance molecular medicine.
Keith says, “What drives my efforts is finding and developing simple solutions for diagnostics testing.”
Peta Owens-Liston, ARUP Science Communications Writer
Related blog:
“Heads in the Cloud: Harnessing the Healing Power of Bioinformatics ”