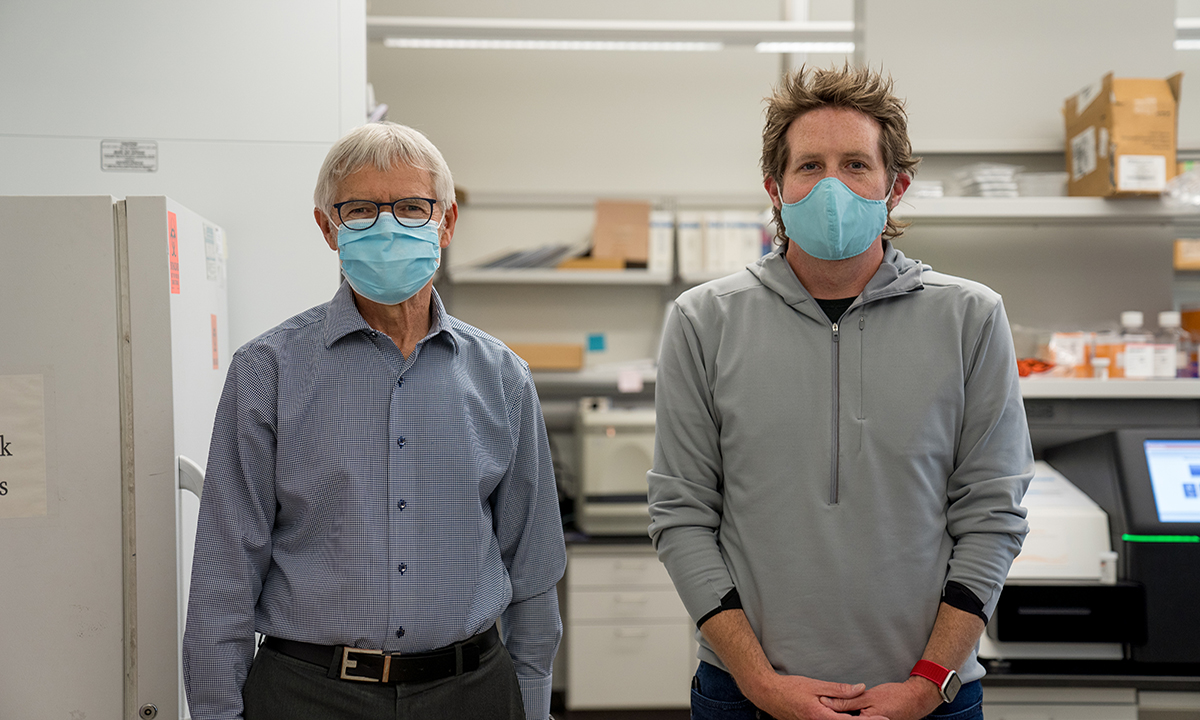
David Hillyard, MD, (left) medical director of ARUP’s Molecular Infectious Diseases Laboratory, and Kelly Oakeson, PhD, (right) Utah Public Health Laboratory chief scientist of bioinformatics and next generation sequencing, are studying the genetics of the COVID-19 virus in Utah.
SALT LAKE CITY — ARUP Laboratories is collaborating with the Utah Department of Health (UDOH) Public Health Laboratory (UPHL) and Boston-based Ginkgo Bioworks to study the genetics of the COVID-19 virus in Utah and analyze how the molecular makeup of the virus is evolving.
The study will help epidemiologists and other healthcare professionals understand how SARS-CoV-2 has spread in Utah and will help them track and prevent future infection, said David Hillyard, MD, medical director of ARUP’s Molecular Infectious Diseases Lab.
“Genetic sequencing and molecular evolutionary analysis of the virus also could also provide important predictive information about the impact of viral evolution on future diagnostic tests, treatments, or vaccines,” said Hillyard, who is collaborating on the study with Kelly Oakeson, PhD, the UPHL’s chief scientist for bioinformatics and next generation sequencing (NGS).
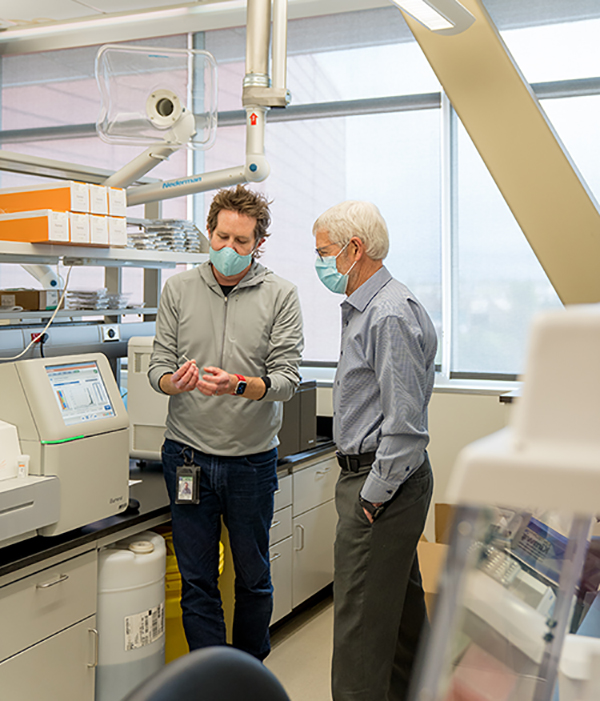

Oakeson and UPHL colleagues rely on genetic sequencing and analysis to help track foodborne pathogens back to their sources — sometimes all the way back to a specific food handler or farm. They also use genetic sequencing to trace the causes of hospital-acquired infections.
“Early in the COVID-19 pandemic, we realized we could apply our expertise to SARS-CoV-2 and potentially track transmission chains back to their sources by obtaining and analyzing the genetic sequences of as many positive COVID-19 specimens as possible,” Oakeson said.
UPHL uses NGS in its analysis. It is a relatively fast technique, but still takes 3-4 days to complete for each sample, Oakeson said.
Because Utah so far has had more than 12,500 confirmed cases of COVID-19, sequencing all of them is more than UPHL can handle alone.
That’s where ARUP comes in. As one of the nation’s four largest reference laboratories, ARUP has expertise in sequencing and sequence analysis, but also the capacity to extract virus from samples in preparation for sequencing by Ginkgo Bioworks, a third partner in the study with extensive NGS capacity, Hillyard said.
“The mathematical methods used to analyze the data require lots of samples to identify accurate and reliable trends, so collaboration is vital to achieve actionable results,” Oakeson said.
The UPHL has already sequenced more than 500 patient specimens and submitted the data to international data repositories to aid in global response — another goal of the project.
With involvement from ARUP and Ginkgo Bioworks, the number of specimens sequenced will increase dramatically, Oakeson said. “The more data we collect, the clearer the picture of SARS-CoV-2 spread in Utah becomes,” he said.
Oakeson hopes that all COVID-19-positive samples in Utah will be analyzed to help determine how COVID-19 has spread in the state and to prevent future transmission.
”Now more than ever, it is important to work together on projects like this to learn more about COVID-19 and how to fight it,” Hillyard said.
About ARUP Laboratories
Founded in 1984, ARUP Laboratories is a leading national reference laboratory and a nonprofit enterprise of the University of Utah and its Department of Pathology. ARUP offers more than 3,000 tests and test combinations, ranging from routine screening tests to esoteric molecular and genetic assays. ARUP serves clients across the United States, including many of the nation’s top university teaching hospitals and children’s hospitals, as well as multihospital groups, major commercial laboratories, group purchasing organizations, military and other government facilities, and major clinics. In addition, ARUP is a worldwide leader in innovative laboratory research and development, led by the efforts of the ARUP Institute for Clinical and Experimental Pathology®. ARUP is ISO 15189 CAP accredited.
Media Contacts
ARUP: Lisa Carricaburu, lisa.carricaburu@aruplab.com, 801-541-5041
Utah Department of Health: Charla Haley, chaley@utah.gov, 801-230-5927