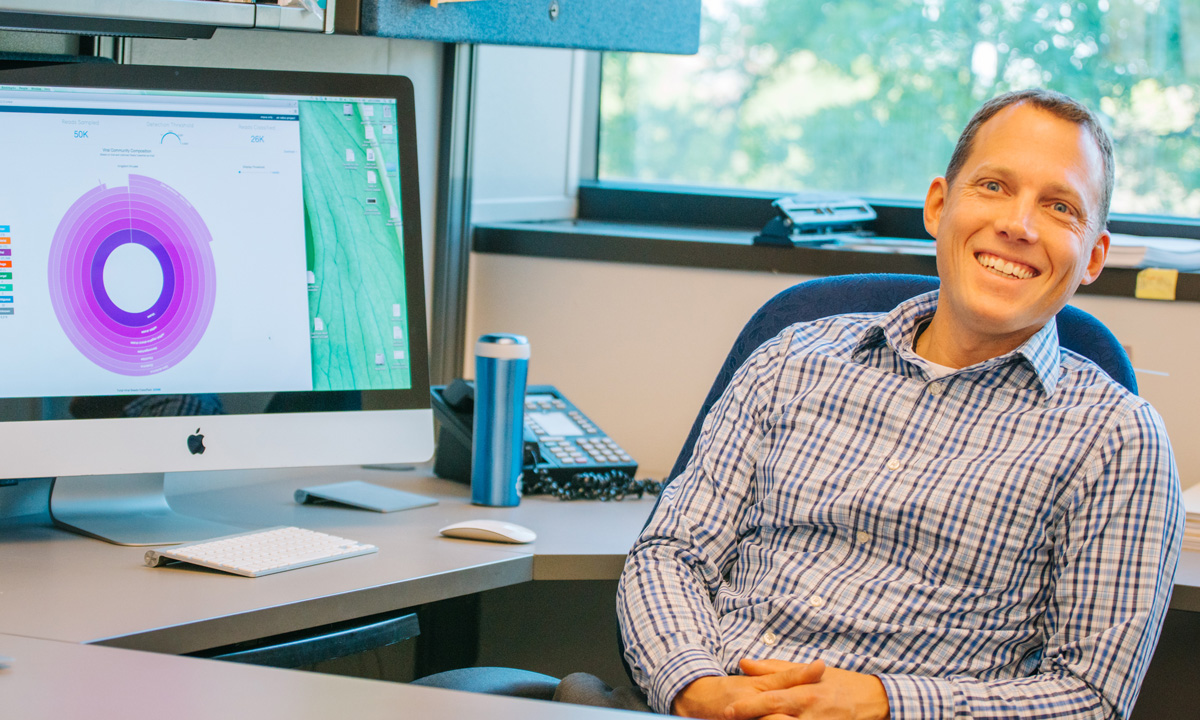
Dr. Robert Schlaberg co-developed Taxonomer, a new kind of pathogen identification tool driven by sophisticated genomic analysis and advanced computational methodologies. This web-based technology quickly and accurately reveals all pathogens in a patient sample.
Scientists at the University of Utah, ARUP Laboratories, and IDbyDNA, Inc., have developed ultra-fast, meta-genomics analysis software called Taxonomer that dramatically improves the accuracy and speed of pathogen detection. In a paper published today in Genome Biology, the collaborators demonstrated the ability of Taxonomer to analyze the sequences of all nucleic acids in a clinical specimen (DNA and RNA) and to detect pathogens, as well as profile the patient’s gene expression, in a matter of minutes.
“Taxonomer can identify an infection without the physician having to decide what to test for, something a PCR-based test cannot do. In other words, a doctor doesn’t have to suspect the cause of a patient’s infection, but can instead simply ask, ‘What does my patient have?’ and Taxonomer will identify the pathogens.”
Robert Schlaberg, MD, Dr Med, MPH
Medical Director, ARUP Laboratories
Infectious diseases are one of the biggest killers in the world. Almost 5 million children under age 5 die each year from infectious diseases worldwide, yet many infections are treatable if the pathogen culprit can be quickly and accurately identified.
“In the realm of infectious diseases, this type of technology could be as significant as sequencing the human genome,” says co-author Mark Yandell, PhD, professor of human genetics at the University of Utah (U of U), H.A. & Edna Benning Presidential Endowed Chair holder, co-director of the USTAR Center for Genetic Discovery, and co-founder of IDbyDNA. “Very few people have inherited genetic disease. But at some point, everyone gets sick from infections.”
It is difficult for infectious pathogens to hide when their genetic material is laid bare. Taxonomer opens up an entirely new approach for infectious disease diagnosis, driven by sophisticated genomic analysis and computational technologies. After a patient’s sample is sequenced, the data are uploaded via the internet to Taxonomer. In less than one minute, the tool displays a thumbnail inventory of all pathogens in the sample, including viruses, bacteria, and fungi. The interactive, real-time user interface of Taxonomer is powered by the iobio system developed by the laboratory of Gabor Marth, DSc, professor of human genetics at the U of U and co-Director of the USTAR Center for Genetic Discovery.
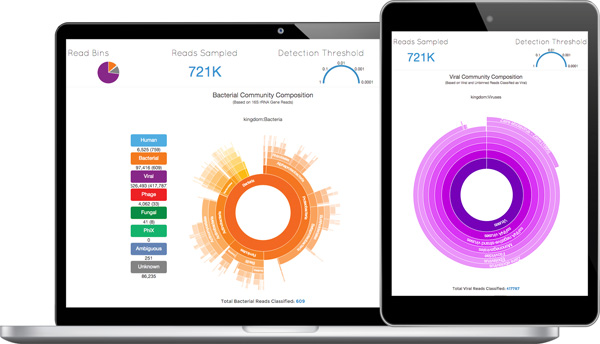
Taxonomer’s interactive display clearly presents the vast genomic data extracted from pathogens found in a patient.
“Our benchmark analyses show Taxonomer being ten to a hundred times faster than similar tools,” says co-author Robert Schlaberg, MD, Dr Med, MPH, a medical director at ARUP Laboratories and cofounder of IDbyDNA. Schlaberg was awarded a $100,000 grant from the Bill and Melinda Gates Foundation to apply Taxonomer toward decreasing high mortality rates of children with infectious diseases in resource-limited settings.
Schlaberg points out that current diagnostic testing still relies heavily on growing cultures of suspected pathogens in the laboratory, which is often inconclusive and time consuming. Even with much faster tests like PCR, the number of pathogens that can be detected is limited.
Schlaberg explains that Taxonomer can identify an infection without the physician having to decide what to test for, something a PCR-based test cannot do. In other words, a doctor doesn’t have to suspect the cause of a patient’s infection, but can instead simply ask, “What does my patient have?” and Taxonomer will identify the pathogens.
In the new study, Taxonomer was put to the test with real-world cases using data published by others and samples provided by ARUP Laboratories and the Centers for Disease Control and Prevention (CDC). Taxonomer determined that some patients who exhibited Ebola-like symptoms in the recent African outbreak did not have Ebola but severe bacterial infections that likely caused their symptoms. “This technology can be applied whenever we don’t know the cause of the disease, including the detection of sudden outbreaks of disease. It is very clear we urgently need more accurate diagnostics to greatly enhance the ability of public health response and clinical care,” says Seema Jain, MD, medical epidemiologist at the CDC.
In the realm of infectious diseases, this type of technology could be as significant as sequencing the human genome. Very few people have inherited genetic disease. But at some point, everyone gets sick from infections.
Mark Yandell, PhD, Professor of Human Genetics, University of Utah
Another unique feature of Taxonomer is its ability to delve into human gene expression profiling, which provides information on how or if the patient’s body is reacting to an infection. “As a clinician, this gives you a better idea, when we identify a pathogen whether it is really the cause of the disease,” says Carrie L. Byington, MD, professor of pediatrics of the U of U and co-director of the Center for Clinical and Translational Science. “This tool will also allow us to determine if the patient is responding to a bacterial or viral infection when we don’t find a pathogen or when we find multiple potential causes.” She says that she sees the exceptional value of this tool for treating children, who experience more life-threatening infections early in life. “Seeing how a host [patient] reacts is extremely valuable; I believe this is a paradigm shift in how we diagnose people. It is why I wanted to be involved.”
In a previous paper published in the Journal of Clinical Microbiology, Schlaberg and his collaborators demonstrated that high-throughput sequencing in combination with Taxonomer can reliably detect pathogens, and identify previously missed pathogens, in patient samples. “Taxonomer provides a critical step forward, as it is extremely fast, accurate, and easy enough to use for implementation in diagnostic laboratories,” says Schlaberg.
Learn more about ARUP Infectious Disease laboratories here.
Peta Owens-Liston, ARUP Science Communications Writer
Related blog
"New Technology Diagnoses Pneumonia Caused by Previously Undetected Pathogens"