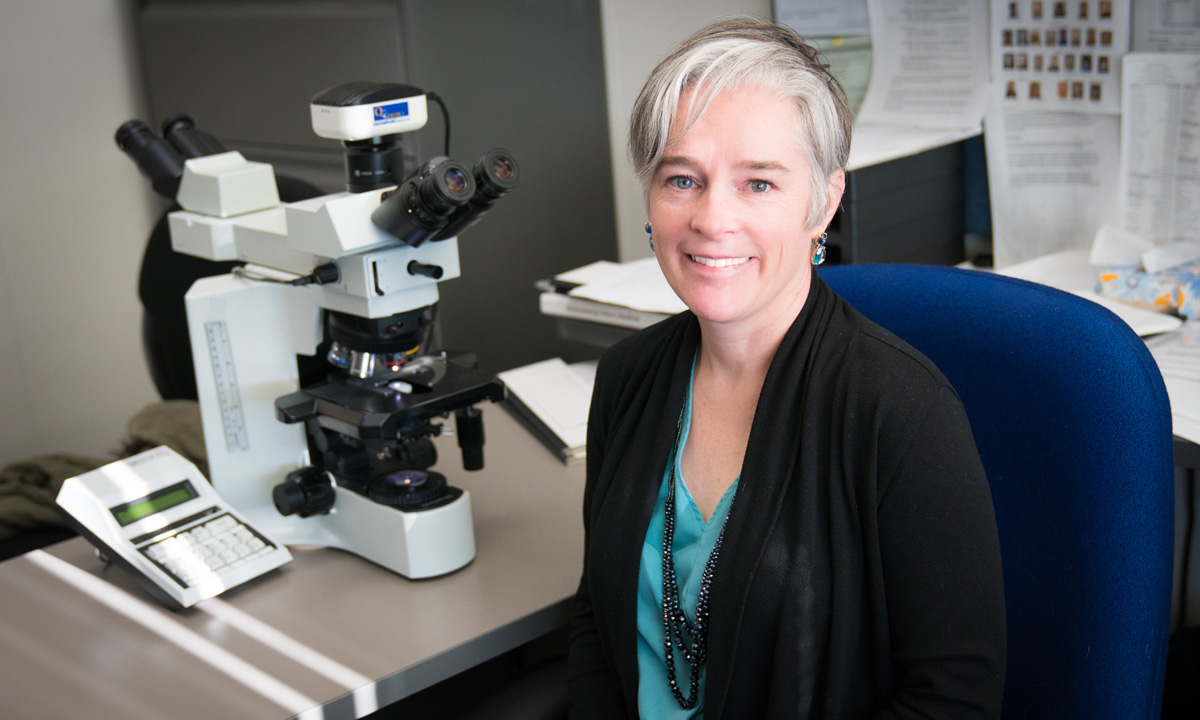
Dr. Kristin Hunt Karner, an ARUP medical director, has current research interests in genetic aspects of myelodysplastic syndrome and other myeloid malignancies. She talks here about NGS for diagnosing myeloid neoplasms.
Kristin Hunt Karner, MD, brings her expertise to ARUP diagnostics every day. A medical director for two ARUP departments, Hematopathology and Molecular Oncology, and an assistant professor of pathology at the University of Utah, she’ll talk here about how the multiple-panel, rapid-throughput system of next-generation sequencing (NGS) genetic material can be used to diagnose myeloid neoplasms, often thought to be precursors to myeloid malignancies (malignancies in tissue) including acute myeloid leukemia (AML). Board certified by the American Board of Pathology in anatomic and clinical pathology, hematopathology, and molecular genetic pathology, Dr. Karner’s areas of focus include lymphoid and myeloid malignancies.
What is a myeloid neoplasm? A malignancy of the cells in the bone marrow of the myeloid (tissue) lineage.
How do you use NGS testing in your own practice?
I use the NGS myeloid malignancy panel every day to assist in my morphologic evaluation of bone marrow biopsies. Such panels are extremely helpful in establishing a malignant diagnosis (or high risk of impending myeloid malignancy), determining important prognostic markers, and tracking certain mutations over time. This type of test also has a high negative predictive value in ruling out myeloid malignancies when a patient presents with cytopenias of unknown etiology.
Dr. Kristin Hunt Karner“These are exciting times, and the way we use the information from this panel is rapidly evolving.”
Medical Director
How has NGS testing for myeloid malignancies changed over the past several years?
This has been an intensive area of study, and we now have a better understanding of the patterns one might see and their associated degrees of risk. For example, while a patient with several mutations might not have a morphologically diagnosable malignancy at the time of biopsy, we now know that the more mutated genes you have and the larger the mutated clone, the more likely that person is to develop a myeloid malignancy. There are also certain genes that have stronger associations with malignancy. So even if these patients don’t have a diagnosable malignancy at the time the biopsy is performed, the presence of these mutations would warrant careful, frequent follow-up in that patient.
What are the pitfalls of NGS test interpretation?
One needs to be aware of the areas that are covered (sequenced) and not covered by the test. In our assay, we are only looking at exonic regions, and in some cases, we don’t look at the entire gene but focus on “hot spots.” It’s also important that the doctor on the receiving end doesn’t assume that the presence of a mutation automatically equals malignancy. There is a condition called “CHIP” (clonal hematopoiesis of indeterminate potential) where elderly individuals who do not have a hematologic malignancy can still possess some of these myeloid mutations, albeit usually only one or two mutations and at low variant allele frequencies. We do know, however, that these people are at higher risk of later developing a hematologic malignancy. We also report tier 2 variants. These are nucleotide variants of uncertain significance that include both potentially pathogenic mutations that have not yet been characterized in the literature, or they could be SNPs (single nucleotide polymorphisms). The tier 2 category implies that we don’t yet have enough information or data to make that call.
When is it appropriate to order NGS myeloid testing?
This test should be ordered by a hematologist after ruling out other causes of the CBC abnormalities (cytoses or cytopenias) and establishing clinical suspicion for a myeloid malignancy, typically supported by abnormal bone marrow morphology. We much more commonly see negative results when the degree of clinical suspicion is lower; this may not be the most cost-effective approach in those cases. This test is not designed to detect common mutations in nonmyeloid malignancies, such as lymphoblastic leukemias, lymphomas, or myelomas, and is of little utility in those cases.
Where do you see this type of testing going in the future? How is it evolving?
The building of massive databases of known mutations and sharing of information is helping us to better identify pathogenic mutations and to distinguish them from benign variants. We are also recognizing patterns that may help in the evolution of our World Health Organization myeloid classification system. In the future, the determination of malignancy may be based more on mutation patterns and frequencies than the current heavy emphasis on morphology. We currently allow for the diagnosis of myelodysplastic syndrome without dysplasia if there is a recurrent, specific cytogenetic abnormality, but we don’t yet allow for this type of diagnosis with molecular mutations. I think this will change in years to come. We may also see less invasive diagnosis and monitoring, requiring only a peripheral blood sample rather than the repeated bone marrow biopsies for patients with certain myeloid malignancies. This may be analogous to the improvements in monitoring for patients with chronic myeloid leukemia, BCR-ABL1 positive, who can be followed with RT-PCR testing on the peripheral blood in most cases. These are exciting times, and the way we use the information from this panel is rapidly evolving.
View ARUP's myeloid malignancies panel by NGS.
Catherine Arnold, Science Communications Writer
Watch “One in a Million,” a short documentary about figuring out one child’s extremely rare disease. This important work is happening at the University of Utah and ARUP.
Related blog
"When Your Own Blood Turns on You: A Journey of Endurance through Multiple Myeloma"